Logistic regression continued
Introduction
Nov 06, 2024
Announcements
- Project: Paper Due November 18th
- Oral R Quiz
📋 AE 19 - Intro to Logistic Regression Continued
- Open up AE 19 and complete Exercise 0.
Logistic regression
Topics
- Use R to fit logistic regression models
- Interpret coefficients of logistic regression models
- Use logistic regression model to calculate predicted odds and probabilities
Computational setup
Data: Framingham Study
This data set is from an ongoing cardiovascular study on residents of the town of Framingham, Massachusetts. We want to use the total cholesterol to predict if a randomly selected adult is high risk for heart disease in the next 10 years.
Variables
- Response:
TenYearCHD:- 1: Patient developed heart disease within 10 years of exam
- 0: Patient did not develop heart disease within 10 years of exam
- Predictors:
totChol: total cholesterol (mg/dL)age: age in years
Logistic regression
Logistic regression model
Logit form: \[\log\big(\frac{\pi}{1-\pi}\big) = \beta_0 + \beta_1~X\]
Probability form:
\[ \pi = \frac{\exp\{\beta_0 + \beta_1~X\}}{1 + \exp\{\beta_0 + \beta_1~X\}} \]
Today: Using R to fit this model.
High risk vs. age
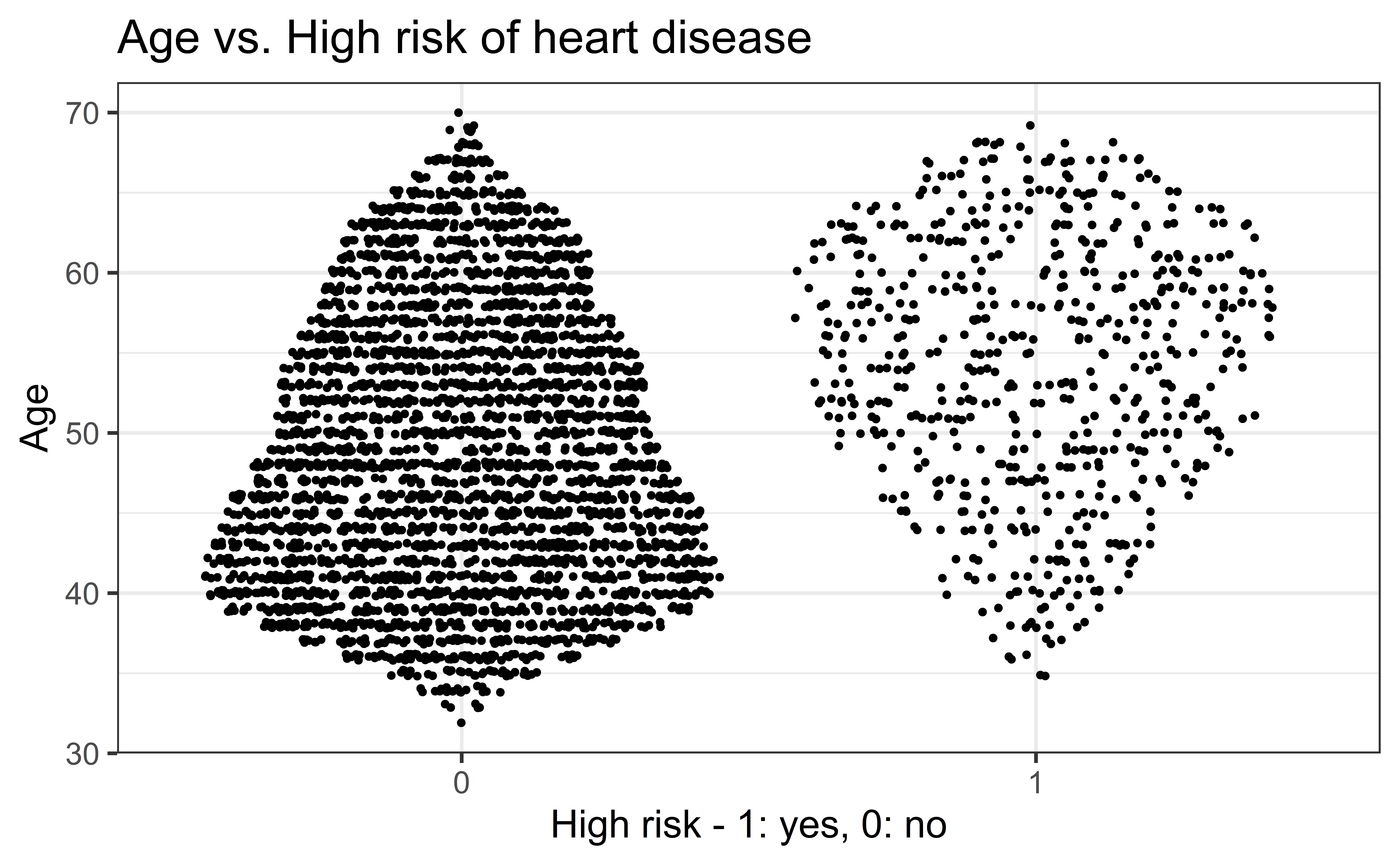
High risk vs. age
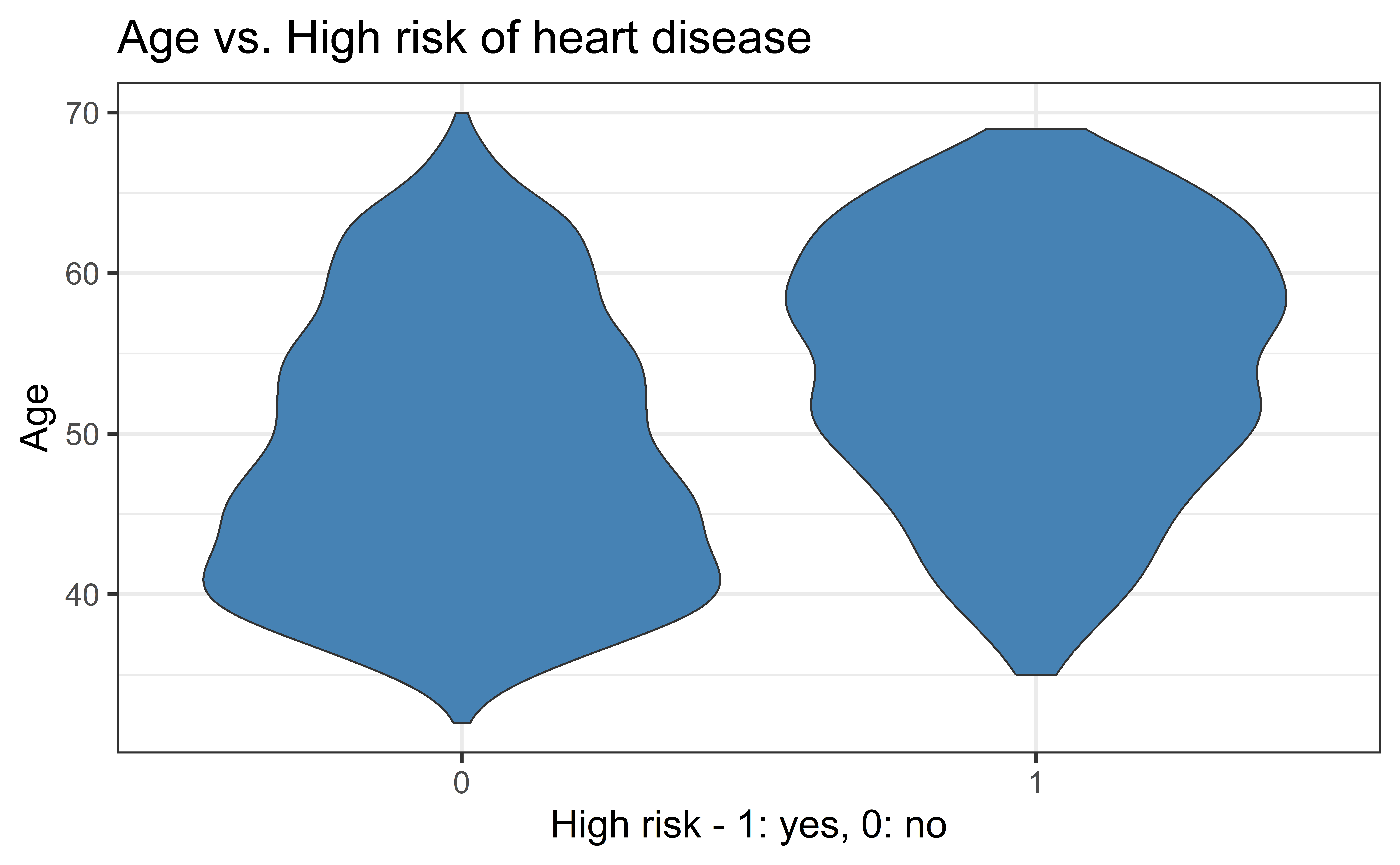
High risk vs. age
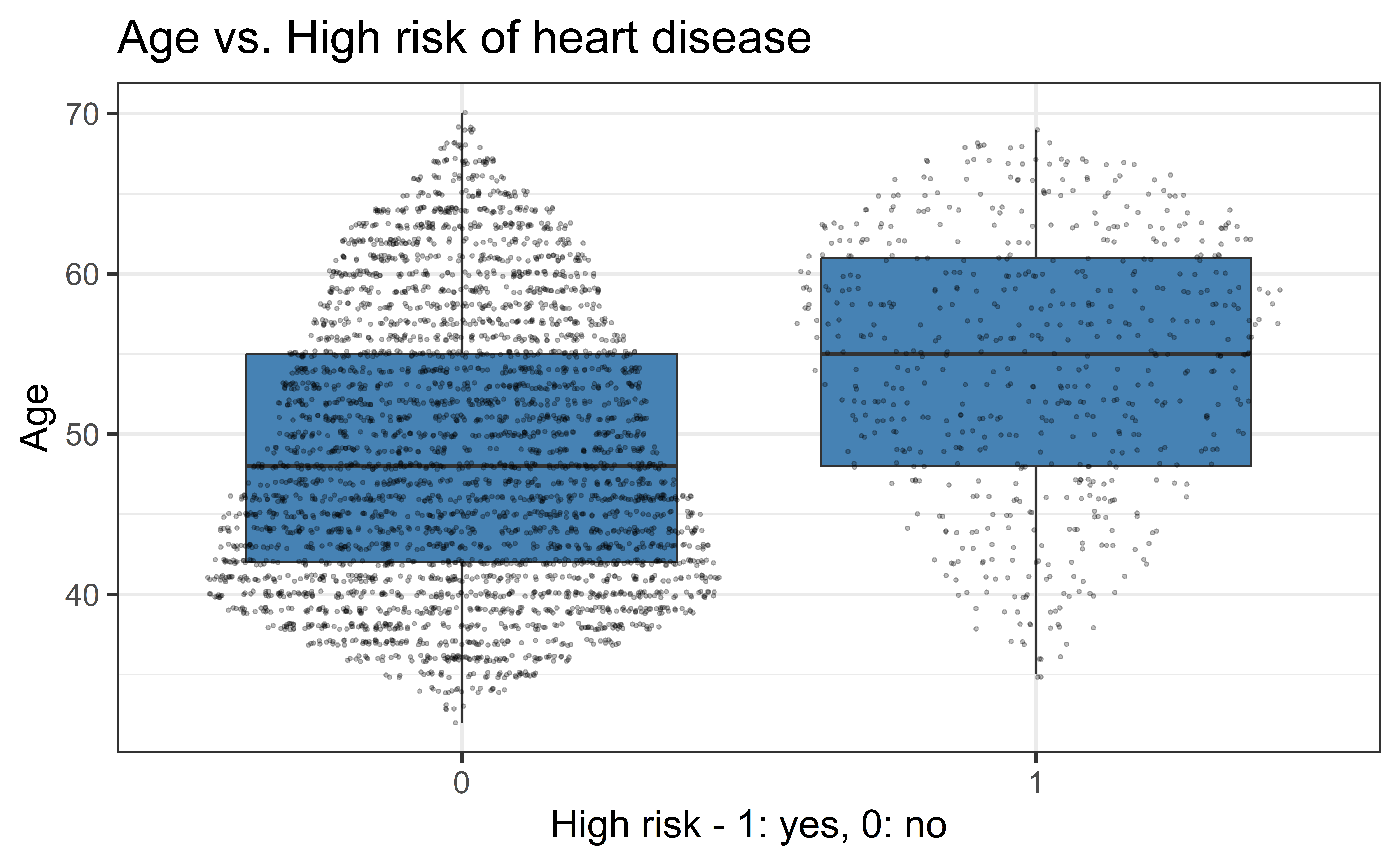
Let’s fit a model
The model
| term | estimate | std.error | statistic | p.value |
|---|---|---|---|---|
| (Intercept) | -5.661 | 0.290 | -19.526 | 0 |
| age | 0.076 | 0.005 | 14.198 | 0 |
\[\textbf{Logit form:}\qquad\log\Big(\frac{\hat{\pi}}{1-\hat{\pi}}\Big) = -5.561 + 0.075 \times \text{age}\]
\[\textbf{Probability form:}\qquad\hat{\pi} = \frac{\exp(-5.561 + 0.075 \times \text{age})}{1+\exp(-5.561 + 0.075 \times \text{age})}\]
where \(\hat{\pi}\) is the predicted probability of developing heart disease in the next 10 years.
Interpreting \(\hat{\beta}\)’s
| term | estimate | std.error | statistic | p.value |
|---|---|---|---|---|
| (Intercept) | -5.661 | 0.290 | -19.526 | 0 |
| age | 0.076 | 0.005 | 14.198 | 0 |
For every addition year of age, the log-odds of developing heart disease in the next 10 years, increases by 0.077.
Complete Exercises 1 - 3.
Interpretability of \(\beta\) for predicted probabilities
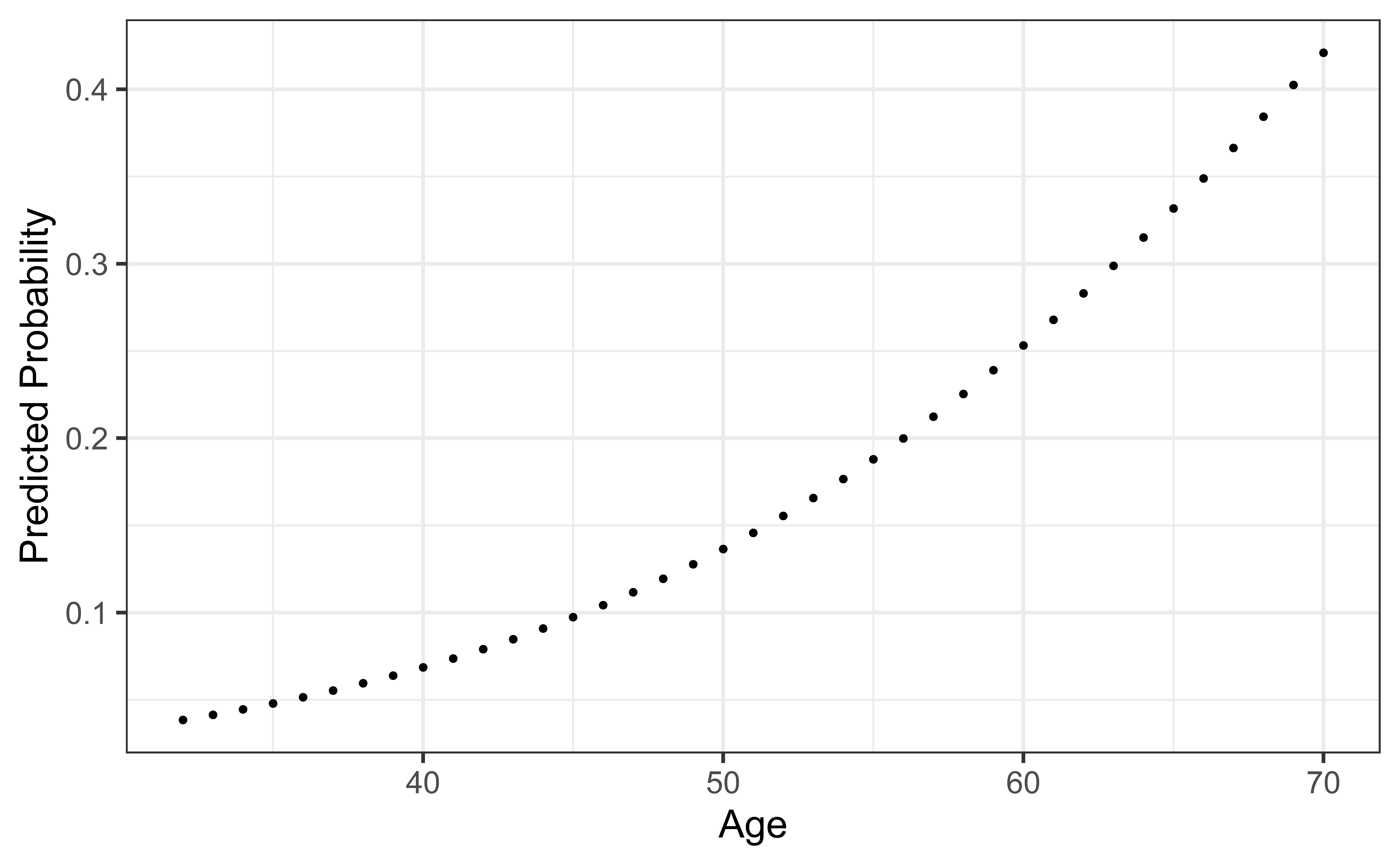
- SLOPE IS CHANGING!
- Increase in \(\hat{\pi}\) due to increase of 1 year of Age depends on what starting age is
glm and augment
The .fitted values in augment correspond to predictions from the logistic form of the model (i.e. the log-odds):
# A tibble: 6 × 8
TenYearCHD age .fitted .resid .hat .sigma .cooksd .std.resid
<dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 0 39 -2.68 -0.363 0.000472 0.891 0.0000161 -0.363
2 0 46 -2.15 -0.469 0.000330 0.891 0.0000192 -0.469
3 0 48 -2.00 -0.504 0.000295 0.891 0.0000200 -0.504
4 1 61 -1.01 1.62 0.000730 0.891 0.000999 1.62
5 0 46 -2.15 -0.469 0.000330 0.891 0.0000192 -0.469
6 0 43 -2.38 -0.421 0.000393 0.891 0.0000182 -0.421Note: The residuals do not make sense here!
For observation 1
\[\text{predicted probability} = \hat{\pi} = \frac{\exp\{-2.650\}}{1 + \exp\{-2.650\}} = 0.066\]
Using predict with glm
Default output is log-odds:
Using predict with glm
More commonly you want the predicted probability:
predict(heart_disease_fit, newdata = heart_disease, type = "response") |> head() |> kable(digits = 3)| x |
|---|
| 0.064 |
| 0.104 |
| 0.119 |
| 0.268 |
| 0.104 |
| 0.085 |
Complete Exercise 4
Recap
- Fit logistic regression models using
glm - Interpreted coefficients in logistic regression models
- Used logistic regression model to calculate predicted odds and probabilities
- Use
predictto make predictions usingglm
