# load packages
library(tidyverse)
library(broom)
library(ggformula)
library(openintro)
library(knitr)
library(kableExtra) # for table embellishments
library(Stat2Data) # for empirical logit
library(countdown)
# set default theme and larger font size for ggplot2
ggplot2::theme_set(ggplot2::theme_bw(base_size = 20))Assessing Logistic Regression Models
Nov 11, 2024
Announcements
- Project: Paper Due November 18th
- Oral R Quiz
📋 AE 20 - Assessing Logistic Regression Models
- Open up AE 20.
Topics
- Checking model conditions for logistic regression
Computational setup
Data
Risk of coronary heart disease
This data set is from an ongoing cardiovascular study on residents of the town of Framingham, Massachusetts. We want to examine the relationship between various health characteristics and the risk of having heart disease.
TenYearCHD:- 1: High risk of having heart disease in next 10 years
- 0: Not high risk of having heart disease in next 10 years
age: Age at exam time (in years)currentSmoker: 0 = nonsmoker, 1 = smoker
Data prep
Conditions
The models
Model 1: Let’s predict TenYearCHD from currentSmoker:
risk_fit <- glm(TenYearCHD ~ currentSmoker,
data = heart_disease, family = "binomial")
tidy(risk_fit, conf.int = TRUE) |>
kable(digits = 3)| term | estimate | std.error | statistic | p.value | conf.low | conf.high |
|---|---|---|---|---|---|---|
| (Intercept) | -1.774 | 0.061 | -28.936 | 0.000 | -1.896 | -1.656 |
| currentSmoker1 | 0.108 | 0.086 | 1.266 | 0.206 | -0.059 | 0.276 |
Model 2: Let’s predict TenYearCHD from age:
Conditions for logistic regression
Linearity: The log-odds have a linear relationship with the predictors.
Randomness: The data were obtained from a random process.
Independence: The observations are independent from one another.
Empirical logit
The empirical logit is the log of the observed odds:
\[ \text{logit}(\hat{p}) = \log\Big(\frac{\hat{p}}{1 - \hat{p}}\Big) = \log\Big(\frac{\# \text{Yes}}{\# \text{No}}\Big) \]
Calculating empirical logit (categorical predictor)
If the predictor is categorical, we can calculate the empirical logit for each level of the predictor.
| currentSmoker | TenYearCHD | n | prop | emp_logit |
|---|---|---|---|---|
| 0 | 1 | 311 | 0.1449883 | -1.774462 |
| 1 | 1 | 333 | 0.1589499 | -1.666062 |
Complete Exercise 2.
Calculating empirical logit (quantitative predictor)
Divide the range of the predictor into intervals with approximately equal number of cases. (If you have enough observations, use 5 - 10 intervals.)
Compute the empirical logit for each interval
You can then calculate the mean value of the predictor in each interval and create a plot of the empirical logit versus the mean value of the predictor in each interval.
Empirical logit plot in R (quantitative predictor)
Created using dplyr and ggplot functions.
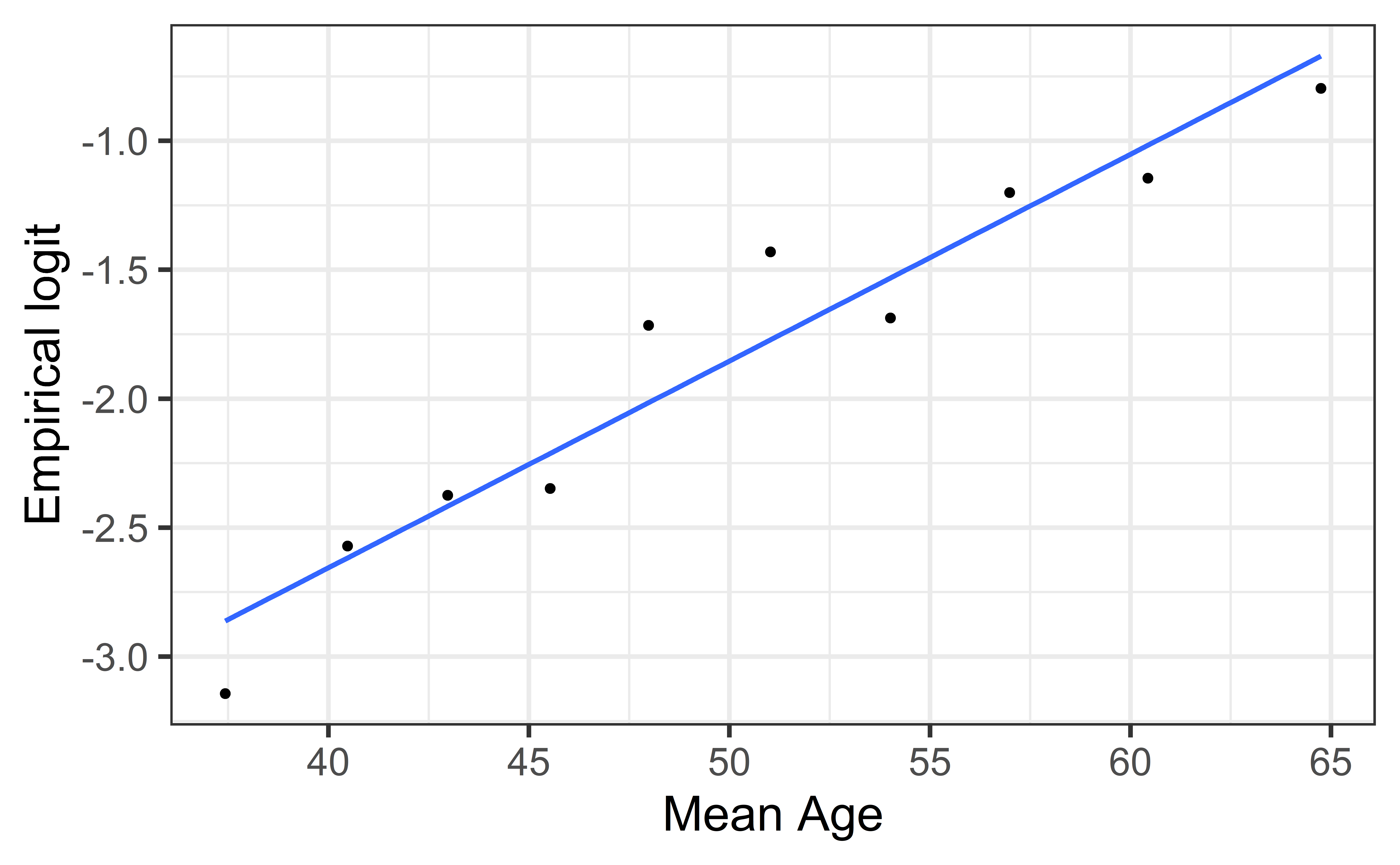
Empirical logit plot in R (quantitative predictor)
Created using dplyr and ggformula functions.
heart_disease |>
mutate(age_bin = cut_interval(age, n = 10)) |>
group_by(age_bin) |>
mutate(mean_age = mean(age)) |>
count(mean_age, TenYearCHD) |>
mutate(prop = n/sum(n)) |>
filter(TenYearCHD == "1") |>
mutate(emp_logit = log(prop/(1-prop))) |>
gf_point(emp_logit ~ mean_age) |>
gf_smooth(method = "lm", se = FALSE) |>
gf_labs(x = "Mean Age",
y = "Empirical logit")Empirical logit plot in R (quantitative predictor)
Using the emplogitplot1 function from the Stat2Data R package
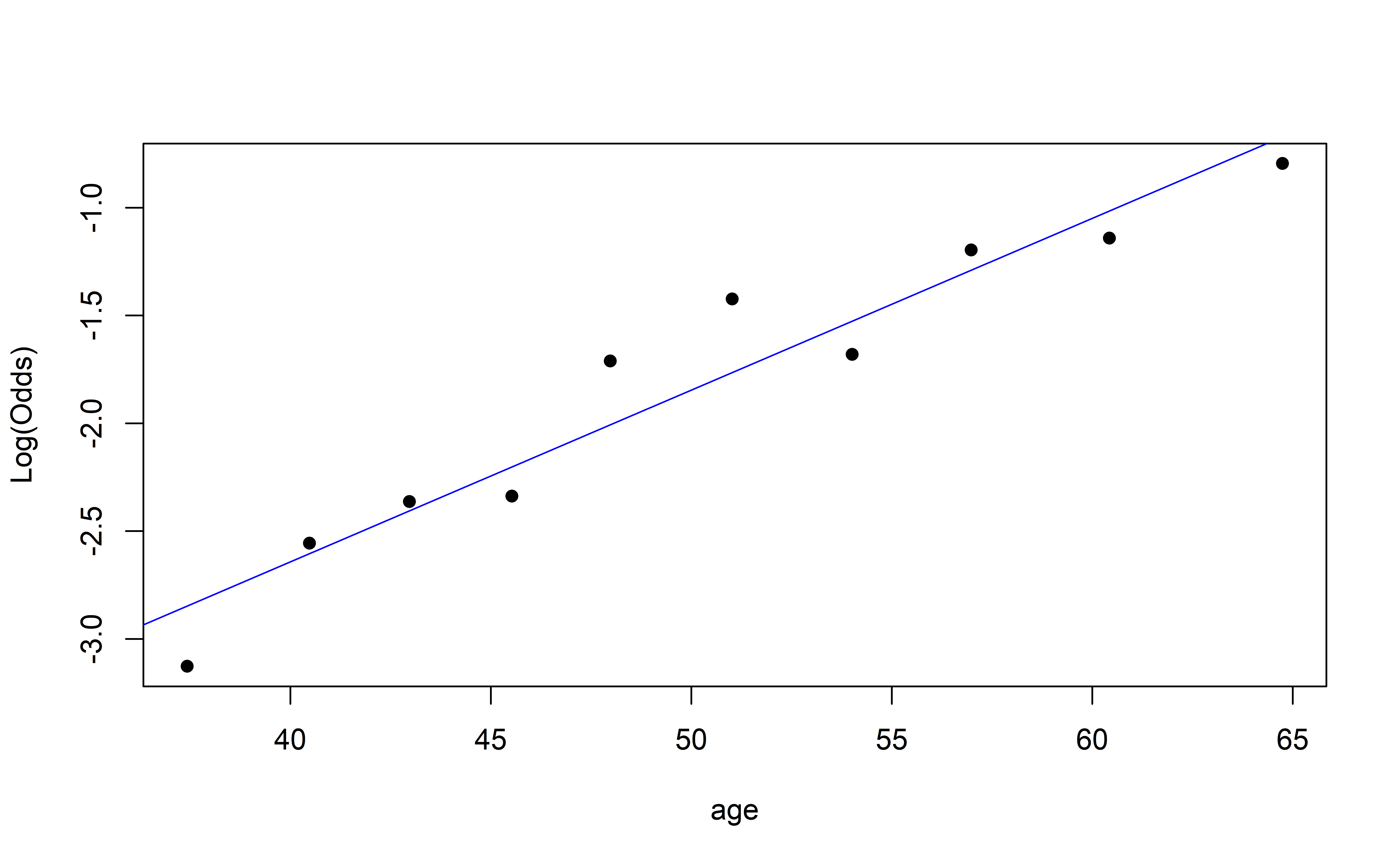
Checking linearity
✅ The linearity condition is satisfied. There is a linear relationship between the empirical logit and the predictor variables.
Complete Exercise 3.
Checking randomness
We can check the randomness condition based on the context of the data and how the observations were collected.
- Was the sample randomly selected?
- If the sample was not randomly selected, ask whether there is reason to believe the observations in the sample differ systematically from the population of interest.
✅ The randomness condition is satisfied. We do not have reason to believe that the participants in this study differ systematically from adults in the U.S. in regards to health characteristics and risk of heart disease.
Checking independence
- We can check the independence condition based on the context of the data and how the observations were collected.
- Independence is most often violated if the data were collected over time or there is a strong spatial relationship between the observations.
✅ The independence condition is satisfied. It is reasonable to conclude that the participants’ health characteristics are independent of one another.
Recap
- Logistic regression conditions
- Linearity
- Randomness
- Independence
Rest of Class
Let’s start thinking about inference in the context of Logistic Regression. Find a spot on the white board with the rest of your group:
- Design a simulation-based hypothesis test to determine whether your predictor is useful. Your answer should include a step by step guide of how to implement the test.
- Design a simulation-based procedure for constructing a confidence interval for the coefficient of your assigned predictor.
