Multiple Logistic Regression
Selection + Transformations
Dec 02, 2024
Announcements
📋 AE 24 - Logistic Regression: Selection and Comparison
- Open up AE 24.
Topics
Investigating interaction terms in logistic regression models
Comparing logistic regression models
Choosing logistic regression models
Transformations in logistic regression
Computational setup
Data
Risk of coronary heart disease
This data set is from an ongoing cardiovascular study on residents of the town of Framingham, Massachusetts. We want to examine the relationship between various health characteristics and the risk of having heart disease.
TenYearCHD:- 1: Developed heart disease in next 10 years
- 0: Did not develop heart disease in next 10 years
age: Age at exam time (in years)education: 1 = Some High School, 2 = High School or GED, 3 = Some College or Vocational School, 4 = College
Data prep
heart_disease <- read_csv("data/framingham.csv") |>
select(TenYearCHD, age, education, male, diaBP) |>
drop_na() |>
mutate(education = as.factor(education),
male = as.factor(male))
heart_disease |> head() |> kable()| TenYearCHD | age | education | male | diaBP |
|---|---|---|---|---|
| 0 | 39 | 4 | 1 | 70 |
| 0 | 46 | 2 | 0 | 81 |
| 0 | 48 | 1 | 1 | 80 |
| 1 | 61 | 3 | 0 | 95 |
| 0 | 46 | 3 | 0 | 84 |
| 0 | 43 | 2 | 0 | 110 |
Interaction Terms
Interactions
Intuitively: There is an interaction between two variables when the influence of one variables on the log-odds depends on what the value of the other variable is.
Recall: Empirical Logit Plot
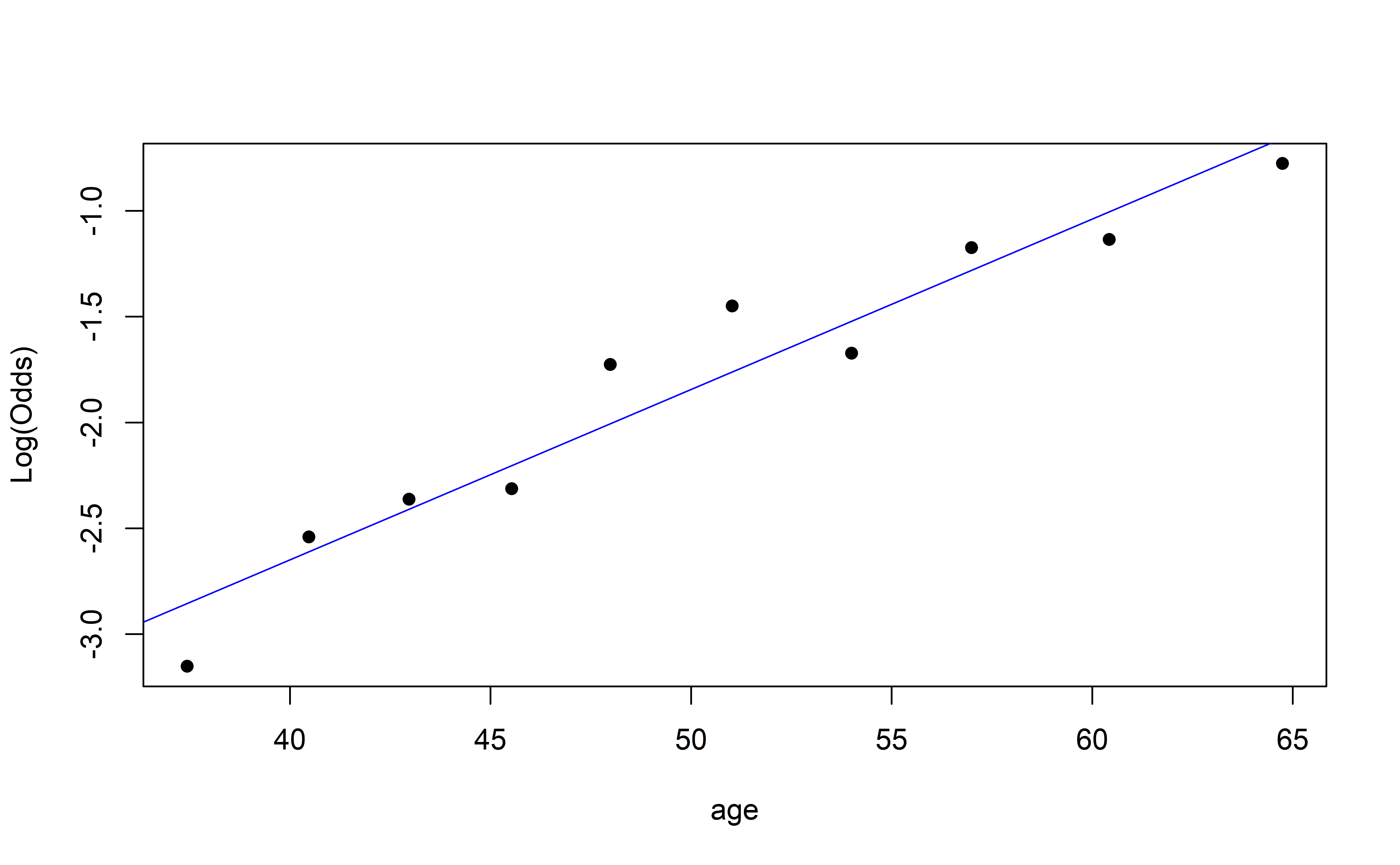
How do we interpret this?
Visualizing Interactions (Quant vs. Cat)
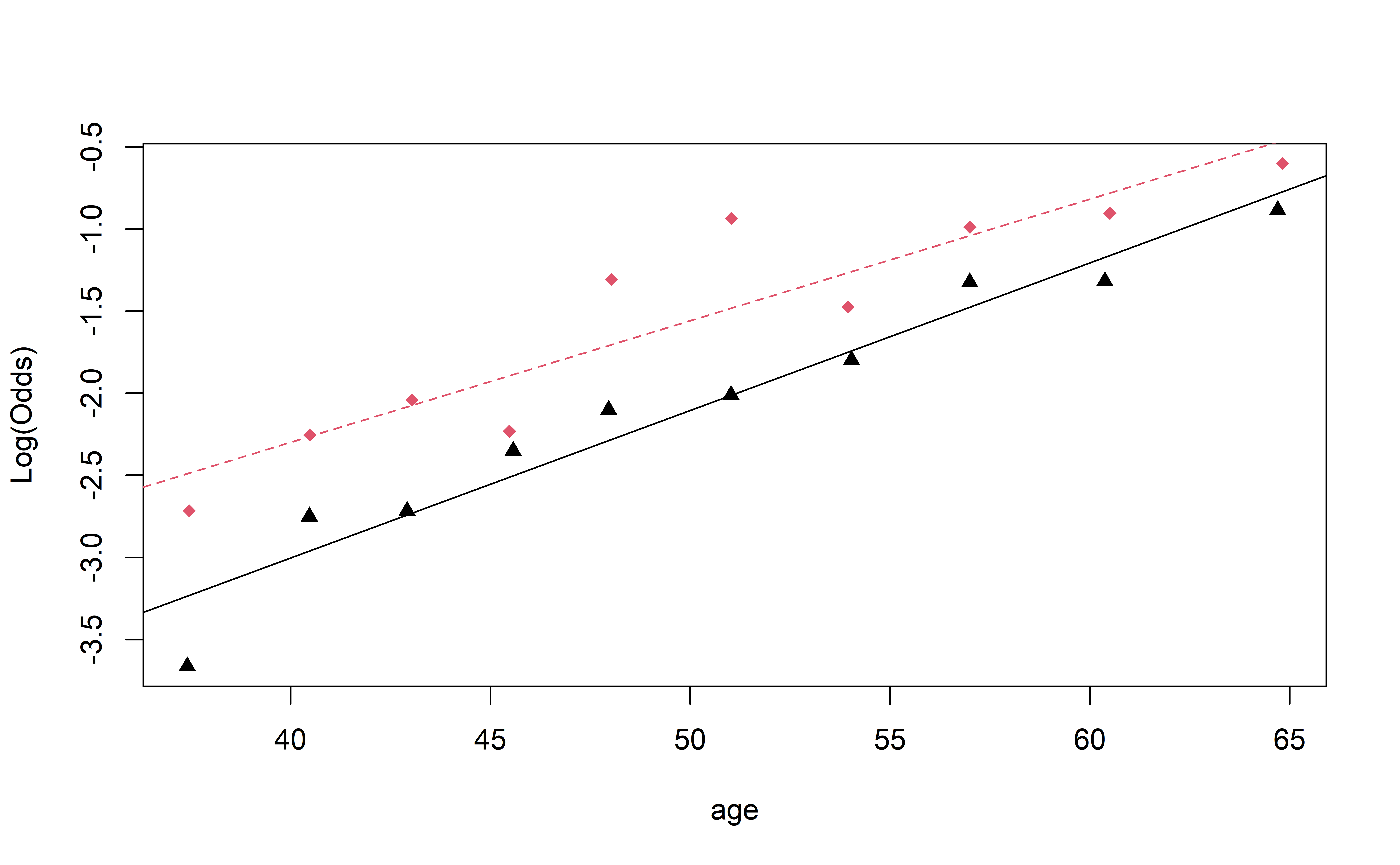
Complete Exercise 2.
Visualizing Interactions (Quant vs. Cat)
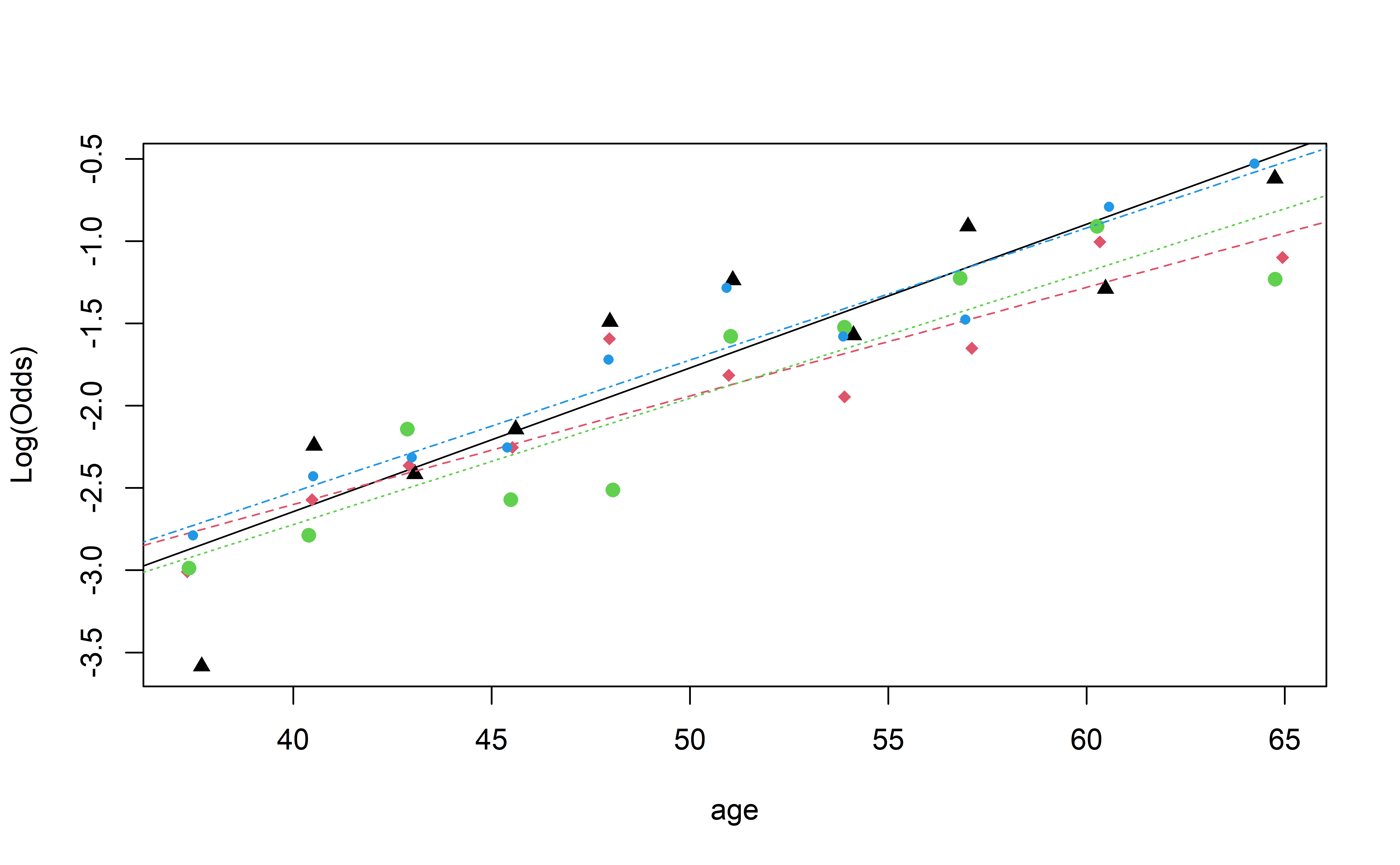
Based on this plot, do you think there is an interaction between age and education?
Models with Interaction
Code
| term | estimate | std.error | statistic | p.value |
|---|---|---|---|---|
| (Intercept) | -6.358 | 0.428 | -14.865 | 0.000 |
| age | 0.085 | 0.008 | 10.883 | 0.000 |
| male1 | 1.308 | 0.583 | 2.245 | 0.025 |
| age:male1 | -0.014 | 0.011 | -1.342 | 0.180 |
Models with Interaction
Two model’s in one!
- Non-male’s: \[\log\left(\frac{\hat{p}}{1-\hat{p}}\right) = -6.358 +0.085\times\text{age}\]
- Male’s: \[ \begin{aligned} \log\left(\frac{\hat{p}}{1-\hat{p}}\right) &= (-6.358+1.308) +(0.085-0.015)\times\text{age}\\ &= -5.05 +0.070\times\text{age} \end{aligned} \]
Complete Exercises 3 and 4.
Model selection
AIC or BIC for model selection
\[ \begin{align} &AIC = - 2 * \log L - {\color{purple}n\log(n)}+ 2(p +1)\\[5pt] &BIC =- 2 * \log L - {\color{purple}n\log(n)} + \log(n)\times(p+1) \end{align} \]
AIC from the glance() function
Let’s look at the AIC for model we fit earlier
Comparing the models using AIC
Let’s compare a few models:
age_model <- glm(TenYearCHD ~ age, data = heart_disease, family = "binomial")
age_male_model <- glm(TenYearCHD ~ age+male, data = heart_disease, family = "binomial")
age_education_model <- glm(TenYearCHD ~ age+education, data = heart_disease, family = "binomial")
age_male_education_model <- glm(TenYearCHD ~ age+ male + education, data = heart_disease, family = "binomial")
age_education_interaction_model <- glm(TenYearCHD ~ age + education + age* education, data = heart_disease, family = "binomial")
all_interaction_model <- glm(TenYearCHD ~ (age + education + male)^2, data = heart_disease, family = "binomial")Comparing the models using AIC
Let’s compare a few models:
aics <- c(glance(age_model)$AIC,
glance(age_male_model)$AIC,
glance(age_male_interaction_model)$AIC,
glance(age_education_model)$AIC,
glance(age_male_education_model)$AIC,
glance(age_education_interaction_model)$AIC,
glance(all_interaction_model)$AIC)
aics[1] 3310.680 3276.964 3277.159 3310.135 3279.342 3314.815 3285.196Based on AIC, which model would you choose? Raise you hand if your AIC is better.
Comparing the models using BIC
Let’s compare the models using BIC
bics <- c(glance(age_model)$BIC,
glance(age_male_model)$BIC,
glance(age_male_interaction_model)$BIC,
glance(age_education_model)$BIC,
glance(age_male_education_model)$BIC,
glance(age_education_interaction_model)$BIC,
glance(all_interaction_model)$BIC)
bics[1] 3323.334 3295.945 3302.468 3341.772 3317.305 3365.433 3367.450Based on BIC, which model would you choose? Raise your hand if your BIC is better.
Choosing a Logistic Regression Model
- Forward/Backward/Stepwise selection all work in the same way
- Can string together a sequence of Likelihood Ratio Tests (but only a few)
Transformations
Power Family of Transformation
Many departures from linearity can be solved with power transformations (e.g. \(X^{power}\))
- For technical reasons, \(power = 0\) corresponds to \(\log\)
Look at empirical log-odds plot
Concave down pattern \(\Rightarrow\) transform down (i.e. \(power < 1\))
- \(\log\) is typically a good first choice
Concave up pattern \(\Rightarrow\) transform up (i.e. \(power > 1\))
Empirical Log-Odds Plot
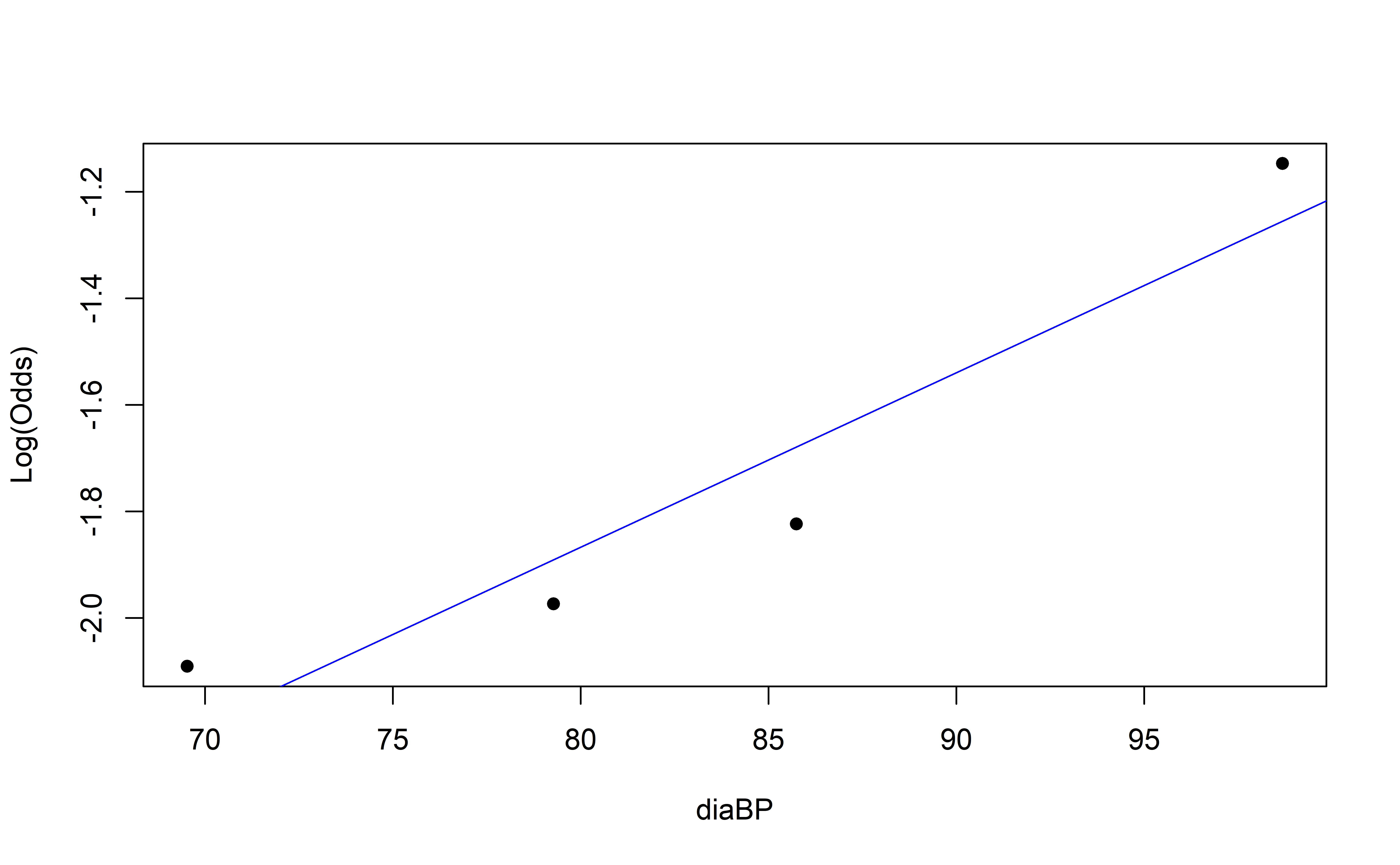
Empirical Log-Odds Plot
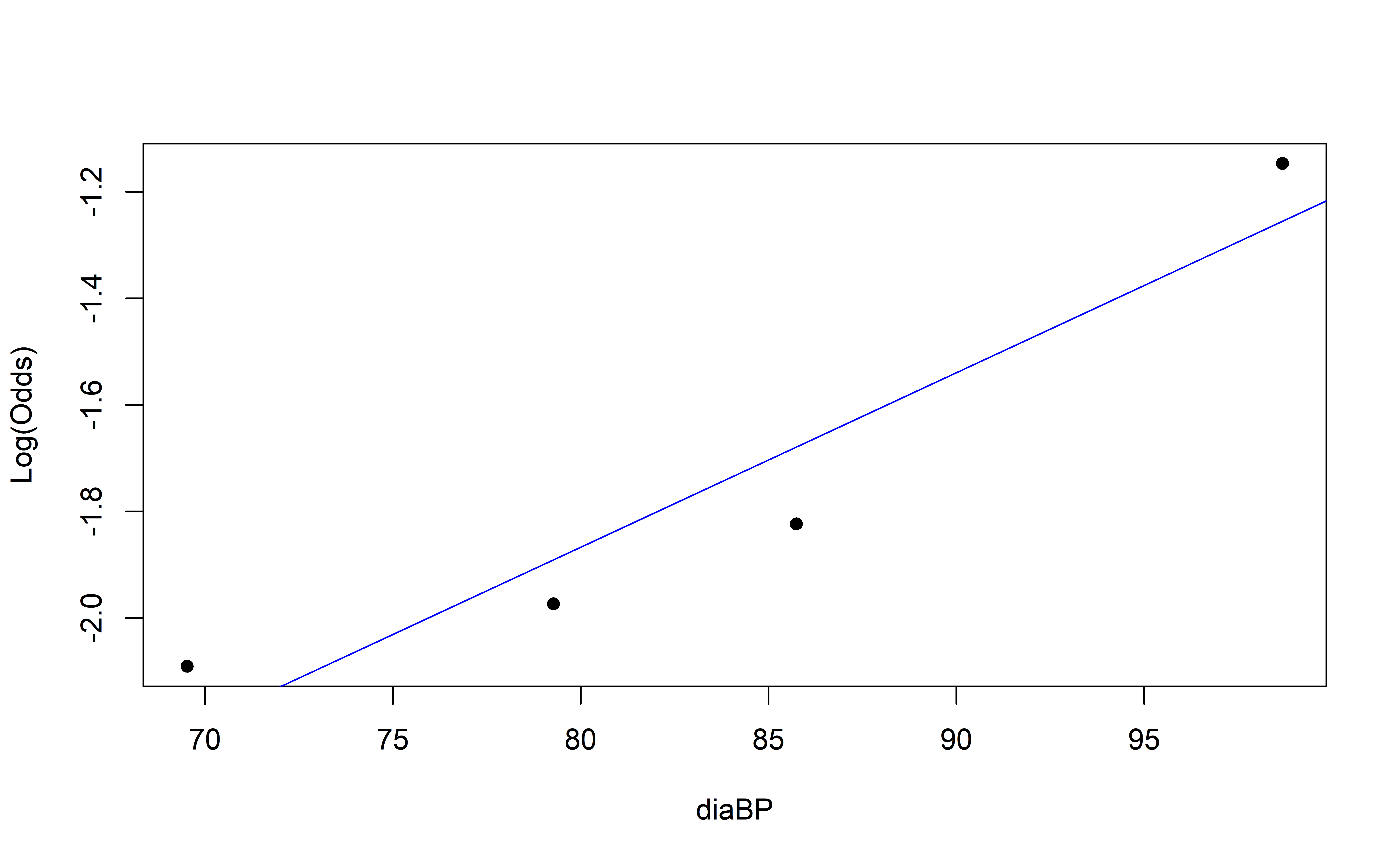
Concave up or down?
Empirical Log-Odds Plot
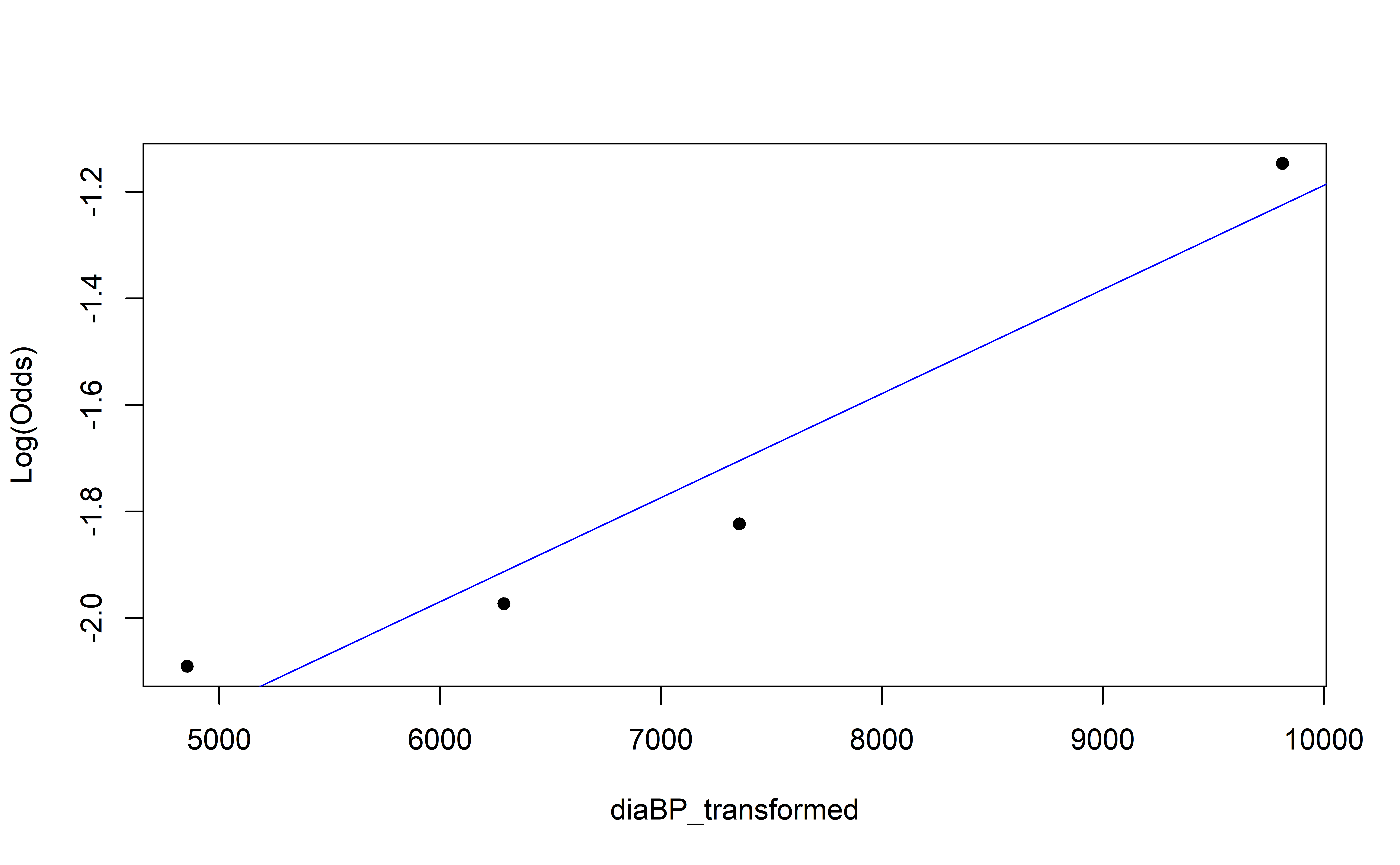
Empirical Log-Odds Plot
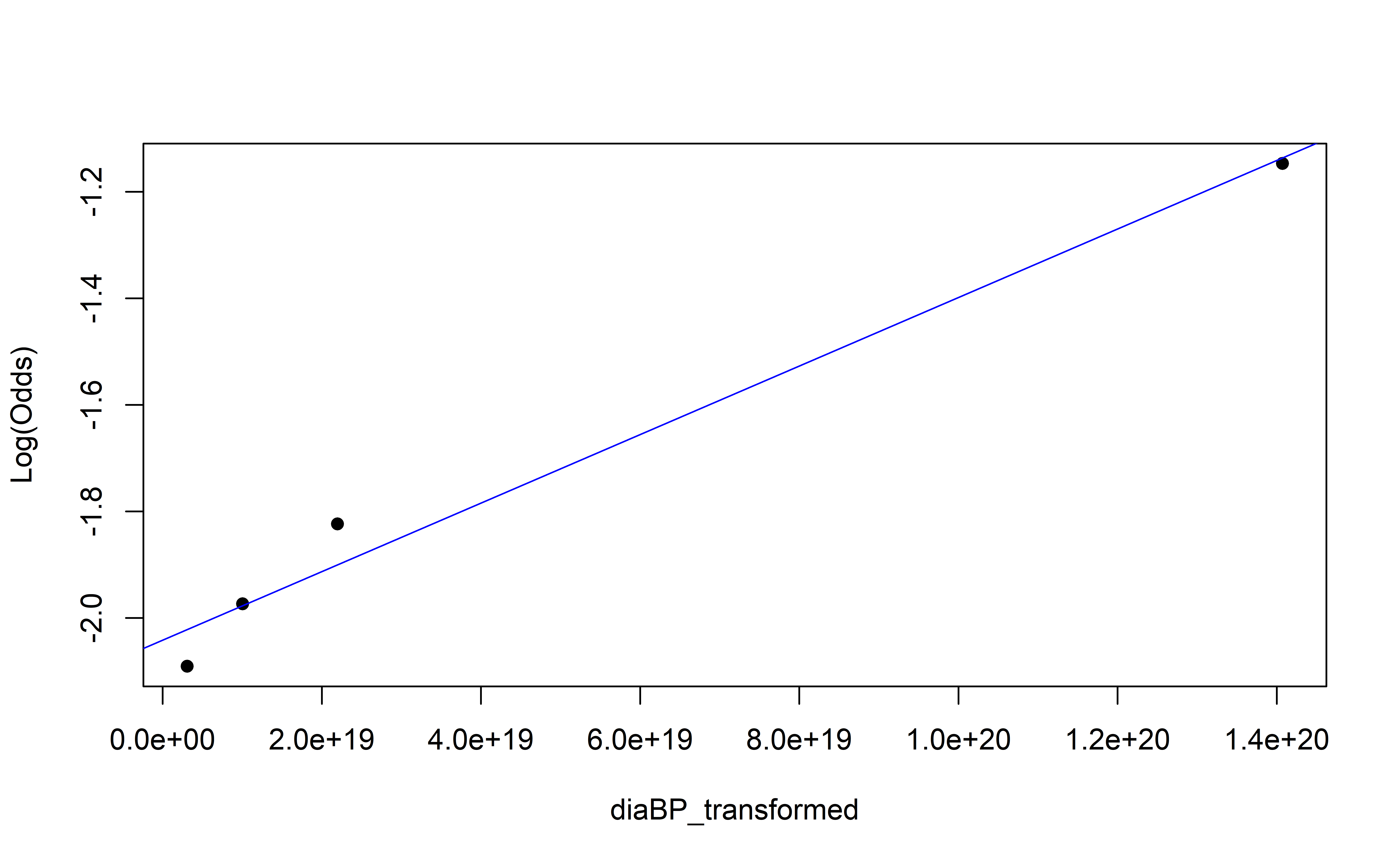
Empirical Log-Odds Plot
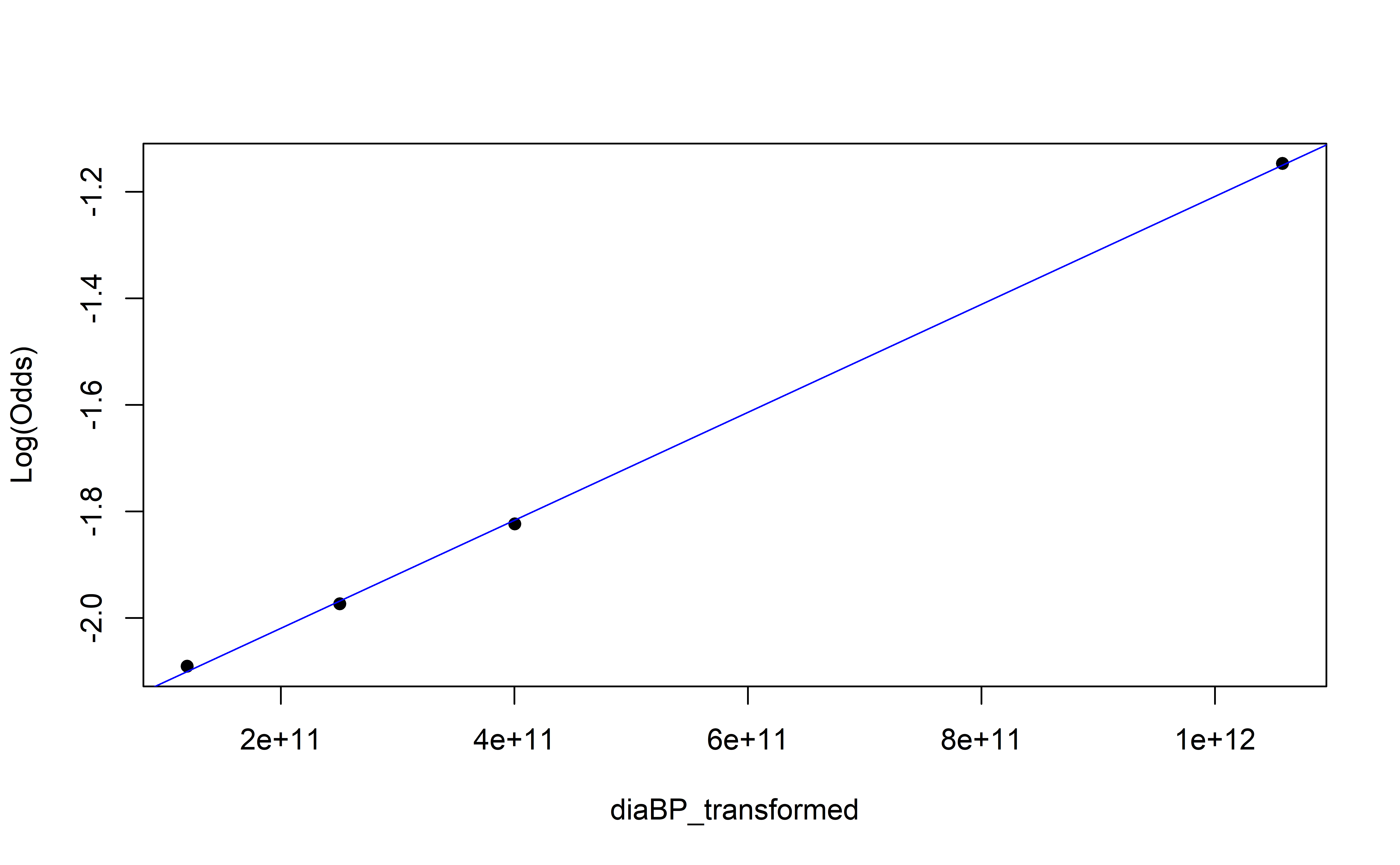
Recap
Interaction terms in logistic regression models
- For quantitative-categorical, creates nested models
Comparing logistic regression models
- AIC/BIC
Choosing logistic regression models
- Same selection techniques as linear regression
Transformations in logistic regression
Can only transform \(X\)
Concave [up/down] \(\Rightarrow\) power-transform [up/down]
